What Is the Transcriptome?
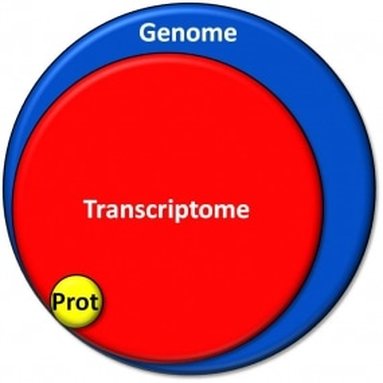
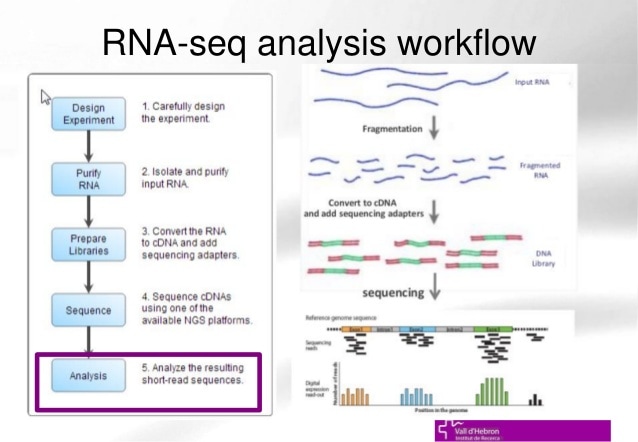
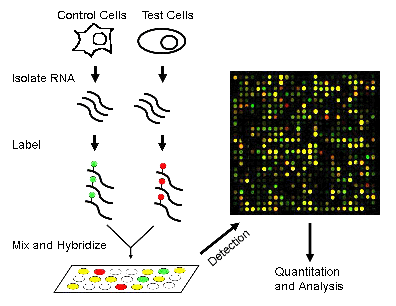
The transcriptome is composed up of all the RNA transcripts in an organism. Transcriptomics is the study of the transcriptome and its function. The goals of transcriptomics is to: determine the transcriptional structure of genes, label all species of transcripts (mRNA, non coding RNA, small RNAs, etc), and to measure changing levels of transcripts under different conditions and during development. RNA-seq and microarrays are popular ways of interpreting RNA expression. The difference between the two is that RNA-seq is specifically a sequence based approach that directly determines the cDNA sequence.
(1).
(1).
conclusions
Looking at mRNA expression is important when analyzing gene expression between people with ADHD and those without. A more characterized gene expression profile of individuals with mutant LPHN3 would help uncover other specific genes that can become differentially expressed in mutant LPHN3. Discovering other genes that are up or down regulated by LPHN3 could possibly lead to new treatments that target crucial interacting genes.
References:
1. Wang, Z., Gerstein, M., & Snyder, M. (2009). RNA-Seq: a revolutionary tool for transcriptomics. Nature Reviews. Genetics, 10(1), 57–63. http://doi.org/10.1038/nrg2484
Images:
Image 1. https://www.dkfz.de/__we_thumbs__/33806_1_ModelTranscriptome.jpg?m=1421890291
figure 1. https://image.slidesharecdn.com/bbr-4-140612021947-phpapp02/95/introduction-to-rnaseq-and-rnaseq-data-analysis-uebuat-bioinformatics-course-session-41-vhir-barcelona-18-638.jpg?cb=1402539761
figure 2. http://www.u.arizona.edu/~gwatts/azcc/flowchart.gif
1. Wang, Z., Gerstein, M., & Snyder, M. (2009). RNA-Seq: a revolutionary tool for transcriptomics. Nature Reviews. Genetics, 10(1), 57–63. http://doi.org/10.1038/nrg2484
Images:
Image 1. https://www.dkfz.de/__we_thumbs__/33806_1_ModelTranscriptome.jpg?m=1421890291
figure 1. https://image.slidesharecdn.com/bbr-4-140612021947-phpapp02/95/introduction-to-rnaseq-and-rnaseq-data-analysis-uebuat-bioinformatics-course-session-41-vhir-barcelona-18-638.jpg?cb=1402539761
figure 2. http://www.u.arizona.edu/~gwatts/azcc/flowchart.gif